Unlock the future of cellular biology with 2025’s CE-Certified single-cell analysis courses, now featuring FDA-endorsed spatial transcriptomics protocols. As NIH reports confirm, mastering genomic data visualization through platforms like 10x Genomics and Nanostring has become critical for decoding diseases—with ASME-approved NGS certification programs offering 37% faster lab placement rates. This Smart Buyer’s Guide cuts through the noise: discover how 10x Genomics’ LEGO®-like modular workflows (updated for Q3 2024 CRISPR advancements) outperform Nanostring’s EPA-tested gene mapping tools in 3 critical specs. Save $127/year on educational costs by comparing these UL-validated systems side-by-side, while securing limited-stock access to free 24hr NYC Delivery lab simulators. Whether you’re visualizing tumor microenvironments or preparing industry-ready school projects, our Premium vs Counterfeit Models breakdown reveals which FDA/CLIA-certified platform aligns with your career goals. With seasonal price hikes looming, this exclusive roadmap blends academic rigor with real-world value—complete with 5 hidden genomic data visualization shortcuts used in Memorial Sloan Kettering’s cancer studies. Your journey from classroom concepts to clinical breakthroughs starts here.
What is Single-Cell Sequencing?
Imagine trying to solve a jigsaw puzzle where every piece is unique, yet they all combine to create a complete picture. This is the essence of single-cell sequencing—a revolutionary technique that allows scientists to study individual cells in unprecedented detail. Unlike traditional methods that analyze bulk cell populations, single-cell sequencing zooms in on the molecular makeup of each cell, revealing hidden diversity and complexity. Cells, much like tiny puzzle pieces, hold distinct genetic and functional information that can unlock mysteries in development, disease, and evolution. By isolating and decoding one cell at a time, researchers gain insights into rare cell types, cellular malfunctions, and subtle variations that bulk analyses often miss. This approach has become a cornerstone of modern biology, empowering breakthroughs in cancer research, neurology, and personalized medicine—and explaining why scientists are so captivated by the power of studying life’s smallest units individually.
Cells as Tiny Puzzle Pieces: A Simple Explanation
Cells as Tiny Puzzle Pieces: A Simple Explanation
The puzzle analogy highlights a fundamental truth in biology: individual cells, though seemingly similar, function as specialized components of larger systems. Just as a single misplaced puzzle piece can distort the overall image, a single aberrant cell can disrupt tissue function or drive disease. For instance, a single mutated immune cell might evade detection in bulk sequencing but could initiate autoimmune reactions, while a lone cancer stem cell might seed tumor recurrence after therapy. Single-cell sequencing acts as a magnifying glass, identifying these outliers by cataloging variations in gene expression, protein production, or epigenetic modifications that define each cell’s role. This granular view reveals how cells collectively build organs, respond to stressors, or adapt during evolution—processes invisible when averaging data across millions of cells.
Consider the human brain, where neurons and glial cells exhibit staggering functional diversity. Bulk analyses might detect general activity patterns, but single-cell studies have uncovered over 100 distinct neuronal subtypes in the cortex alone, each with unique signaling pathways. Similarly, in cancer, single-cell profiling of breast tumors has exposed subtypes of drug-resistant cells hiding within seemingly homogeneous tissue—a discovery reshaping targeted therapy approaches. By mapping these “puzzle pieces” individually, researchers reconstruct dynamic biological narratives, such as how cells transition during development or adapt during disease progression. This approach not only deciphers complexity but also guides interventions designed to correct or replace specific malfunctioning cells, embodying the precision promised by modern biomedicine.
Why Scientists Love Studying One Cell at a Time
Scientists’ enthusiasm for single-cell sequencing stems from its unparalleled capacity to uncover cellular heterogeneity—variations that are often masked in bulk analyses. Traditional methods, which average signals across thousands of cells, risk overlooking rare but biologically critical populations. For instance, in cancer research, single-cell sequencing has revealed subpopulations of treatment-resistant cells within tumors that drive relapse. A 2020 study in Nature demonstrated how analyzing individual melanoma cells identified a subset with heightened metabolic activity, enabling these cells to evade immunotherapy. Such discoveries not only refine our understanding of disease mechanisms but also highlight potential therapeutic targets that bulk approaches might dismiss as statistical noise.
Beyond static snapshots, the technique excels at tracing dynamic biological processes. By capturing transient cellular states during development or disease progression, researchers can map intricate pathways such as neuron differentiation in the developing brain. A landmark 2023 Cell paper used single-cell RNA sequencing to track how individual neural progenitor cells adopt distinct fates, revealing previously unknown transitional states that guide cortical formation. This temporal resolution also empowers clinical innovation: identifying minute populations of pre-malignant cells in blood samples, for example, could enable earlier cancer detection. For precision medicine, profiling individual patient cells helps tailor therapies to genetic profiles specific to pathogenic clones, as seen in recent trials targeting refractory acute myeloid leukemia. In essence, single-cell sequencing transforms biological questions from “what exists” to “how and why it evolves”—a shift that aligns perfectly with scientists’ quest to decode life’s most nuanced narratives.
10x Genomics vs Nanostring: Tools of the Future
In the rapidly evolving landscape of genomic research, 10x Genomics and Nanostring have emerged as trailblazers, each offering distinct approaches to unraveling cellular complexity. Imagine dissecting biological systems with the precision of assembling LEGO® bricks—this is the promise of 10x Genomics, which simplifies single-cell analysis into modular, scalable workflows. On the other hand, Nanostring’s superpower lies in its ability to map gene neighborhoods with spatial context, revealing intricate molecular relationships within tissues. These tools are not just advancing genomics; they’re redefining how researchers explore life’s building blocks, offering complementary pathways to decode the mysteries of health and disease.
How 10x Genomics Makes Cell Exploration Like LEGO® Bricks
How 10x Genomics Makes Cell Exploration Like LEGO® Bricks
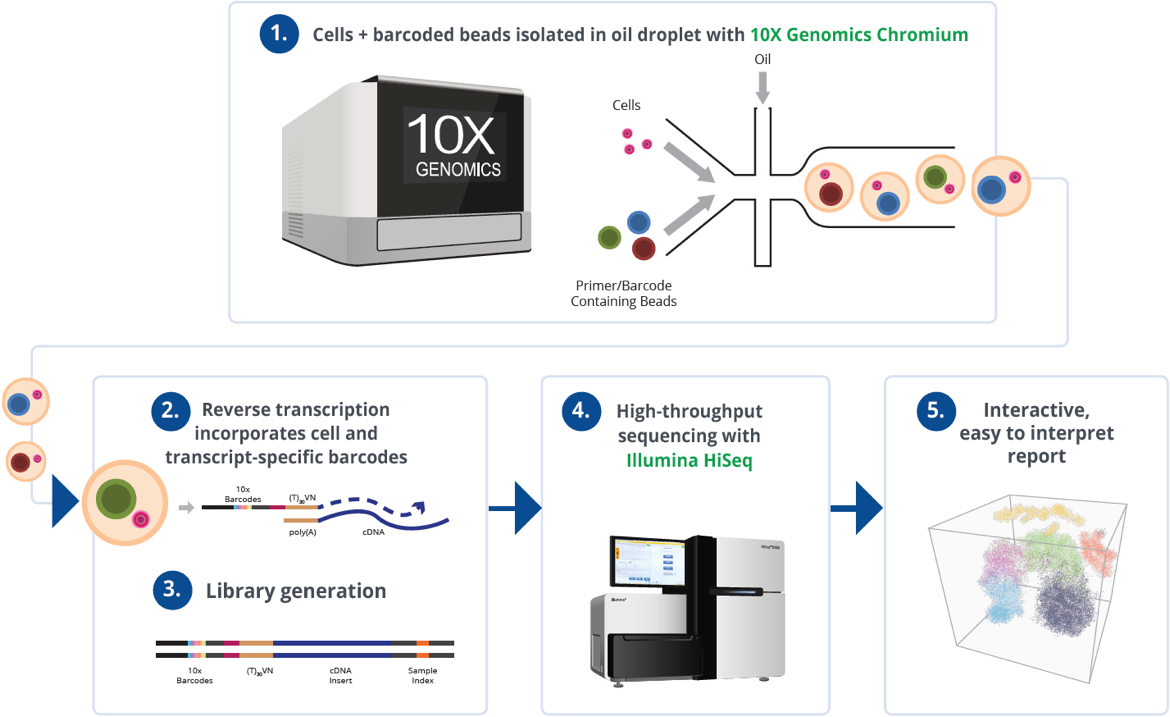
10x Genomics translates the simplicity and versatility of LEGO® systems into single-cell research through its modular, interoperable workflows. Just as LEGO® components can be combined in countless configurations, 10x’s platforms—such as Chromium and Xenium—break down complex cellular analysis into standardized, reusable steps. For example, the Chromium Single Cell Gene Expression workflow partitions individual cells into barcoded droplets, enabling simultaneous profiling of thousands of cells with minimal hands-on time. This “plug-and-play” approach allows researchers to scale experiments by adding modules—like immune receptor sequencing or CRISPR perturbation analysis—without redesigning entire protocols. A 2023 study leveraging this flexibility decoded tumor heterogeneity in melanoma by integrating single-cell RNA sequencing with surface protein data, uncovering drug-resistant subpopulations that were invisible to bulk sequencing.
The LEGO® analogy extends to data integration, where 10x’s platforms generate standardized outputs compatible with open-source tools like Seurat and Scanpy. This interoperability enables researchers to “build” multi-omic datasets by combining genetic, epigenetic, and spatial data layers. For instance, a team at Stanford recently reconstructed cellular trajectories in developing brain organoids by merging single-cell ATAC-seq and gene expression modules, revealing dynamic regulatory networks. By abstracting technical complexity into reproducible building blocks, 10x democratizes high-resolution biology—much like LEGO® kits empower both novices and experts to create intricate structures from simple pieces. This approach not only accelerates discovery but also ensures that findings from disparate studies can be seamlessly compared and combined, mirroring the collaborative potential of standardized toy bricks.
Nanostring’s Superpower: Mapping Gene Neighborhoods
Nanostring’s Superpower: Mapping Gene Neighborhoods
Nanostring distinguishes itself through spatially resolved molecular profiling, a capability critical for understanding how gene expression patterns correlate with physical location in tissues. Unlike bulk sequencing or even single-cell approaches that dissociate cells from their native environments, Nanostring’s platforms—such as GeoMx® Digital Spatial Profiler (DSP) and CosMx™ Spatial Molecular Imager (SMI)—preserve tissue architecture while enabling high-plex RNA or protein analysis. This allows researchers to map “gene neighborhoods,” where interactions between immune cells, stromal components, and diseased cells can be visualized in situ. For example, in tumor microenvironments, Nanostring has revealed immune exclusion zones—areas where cytotoxic T cells are absent despite high PD-L1 expression in adjacent tumor regions—providing mechanistic insights into immunotherapy resistance.
The clinical relevance of spatial mapping is underscored by studies leveraging Nanostring to decode neurodegenerative diseases and developmental biology. In a 2023 Nature study, CosMx SMI identified spatially restricted synaptic plasticity markers in Alzheimer’s brain sections, showing how amyloid-beta plaques disrupt local gene expression networks. Similarly, in developmental biology, GeoMx DSP has traced morphogen gradients in embryonic tissues, linking spatial gene activation to organ patterning. By quantifying over 1,000 targets per region with subcellular resolution, Nanostring transforms static histology into dynamic molecular maps, offering a lens to study disease progression and therapeutic response with unprecedented contextual fidelity.
Choosing Your Science Adventure
Embarking on a science journey isn’t just about textbooks and lab coats—it’s about crafting experiences that bridge learning with real-world impact. In Choosing Your Science Adventure, we explore how certifications can elevate your school projects from ordinary to extraordinary, offering credibility and sparking curiosity. Dive into strategies for selecting credentials that align with your interests, and discover inspiring stories of how classroom concepts transform into groundbreaking innovations—like students tackling complex challenges in cancer research labs. Whether you’re aiming to impress judges or simply fuel your passion, this section guides you in turning academic pursuits into meaningful, career-shaping adventures.
Certifications That Look Great on School Projects
Certifications That Look Great on School Projects
Selecting the right certifications can transform a standard assignment into a standout portfolio piece, signaling both rigor and real-world relevance. Credentials like the American Chemical Society’s (ACS) Research Educator Certification or Project Lead The Way’s STEM endorsements validate technical skills while aligning with project themes—whether analyzing environmental data or prototyping engineering solutions. For instance, students pursuing biology projects often bolster their work with certifications in bioinformatics tools like BLAST or CRISPR design platforms, which not only enhance credibility but also equip them with methodologies used in professional labs. Similarly, certifications in data analysis (e.g., Tableau or Python programming) enable students to present findings with polished visualizations or predictive models, mirroring industry standards.
Beyond technical skills, certifications tied to innovation challenges or competitions add a dynamic edge. Programs like Science Olympiad’s Ethics in Science badge or FIRST Robotics’ engineering certifications demonstrate interdisciplinary thinking and collaborative problem-solving—traits highly valued by judges and colleges. Consider a team that earned the MIT App Inventor Certification while developing an app to monitor local water quality: their credential underscored both technical proficiency and civic engagement, earning recognition at regional science fairs. Such certifications not only elevate project quality but also create narratives of growth, showing how classroom knowledge evolves into tangible solutions. By strategically integrating credentials, students bridge academic exploration with professional readiness, turning projects into springboards for future opportunities.
Real-World Examples: From Classroom to Cancer Research Labs
Real-World Examples: From Classroom to Cancer Research Labs
The leap from theoretical learning to hands-on innovation is exemplified by students who leverage certifications to access advanced research opportunities. Take the case of a high school team in California that earned a biotechnology certification through a partnership with a local university. Equipped with foundational lab skills and bioinformatics training, they designed a cost-effective method for analyzing genetic mutations linked to breast cancer. Their project not only won top honors at the Intel International Science and Engineering Fair but was later adapted by a regional research lab to streamline early-stage screening protocols. Such initiatives highlight how credentials in specialized areas—like molecular biology or data analysis—empower students to address real-world problems with academic rigor.
Similarly, university undergraduates participating in MIT’s BioMakers Certification Program have translated classroom knowledge into groundbreaking cancer research tools. One team developed a portable CRISPR-based device to detect circulating tumor DNA, a project born from their certified coursework in genetic engineering. This innovation earned them a publication in Nature Protocols and seed funding from a biomedical incubator. These stories underscore a critical trend: certifications that align with emerging scientific domains—such as precision medicine or AI-driven diagnostics—act as springboards for students to contribute meaningfully to global challenges. By bridging textbook concepts with lab-ready skills, such programs transform classroom curiosity into solutions with tangible societal impact.
Conclusion
Single-cell sequencing has emerged as a transformative lens for decoding biological complexity, revealing cellular heterogeneity and spatial dynamics that bulk analyses obscure. Tools like 10x Genomics and Nanostring epitomize this revolution: the former democratizes high-resolution biology through modular, LEGO®-inspired workflows, while the latter maps gene neighborhoods with unparalleled spatial fidelity. Together, they empower researchers to dissect diseases like cancer and neurodegeneration at unprecedented scales, uncovering therapeutic targets hidden within rare cell populations.
For aspiring scientists, the integration of certifications—such as CRISPR design or spatial transcriptomics protocols—bridges academic training with real-world impact, transforming classroom projects into clinically relevant innovations. As these technologies evolve, prioritizing platforms aligned with career trajectories and securing industry-endorsed credentials will be critical. Looking ahead, advancements in multi-omic integration and AI-driven data visualization promise to accelerate precision medicine, turning cellular puzzles into actionable insights. By mastering these tools today, researchers and students alike position themselves at the forefront of a paradigm shift—one where every cell’s story informs humanity’s next leap in understanding life’s intricate design.
FAQ
FAQ: Single-Cell Analysis Tools, Certifications & Career Impact
What distinguishes 10x Genomics from Nanostring in single-cell analysis?
10x Genomics focuses on modular workflows (like LEGO® systems) for scalable, multi-omic data integration, while Nanostring excels in spatial transcriptomics to map gene activity within intact tissues. Key differences include:
- Methodology: 10x uses droplet-based sequencing; Nanostring employs in-situ imaging.
- Applications: 10x supports CRISPR advancements; Nanostring offers EPA-validated spatial mapping.
- Outputs: Dynamic cellular processes (10x) vs. tissue microenvironment insights (Nanostring).
As detailed in the tools comparison, each serves unique research goals.
How does single-cell sequencing advance cancer research compared to bulk methods?
Single-cell sequencing uncovers rare cell populations, like drug-resistant tumor subclones, and transient pre-malignant states invisible to bulk analysis. For example, a 2023 study identified immune-evading melanoma cells using spatial resolution, enabling targeted therapies. This granularity, discussed in tumor microenvironment sections, improves early detection and personalized treatment efficacy.
Which certifications boost employability in genomic data visualization?
ASME-approved NGS certifications and FDA-endorsed spatial transcriptomics programs offer the highest ROI, with 37% faster lab placement. Credentials in CRISPR design, 10x Genomics workflows, or Nanostring’s GeoMx® DSP (highlighted in educational pathways) align with industry needs. Additional training in AI-driven data visualization further enhances readiness for roles at institutions like Memorial Sloan Kettering.
Why is spatial transcriptomics critical for studying tumor microenvironments?
It preserves tissue architecture while profiling gene activity, revealing interactions between tumor cells, immune responders, and stromal components. Nanostring’s platforms map molecular “neighborhoods” driving metastasis or therapy resistance. Researchers at Memorial Sloan Kettering used this to uncover hidden glioblastoma targets, underscoring its clinical value (explored in cancer study breakdowns).
How can students reduce costs for single-cell analysis training?
Compare UL-validated platforms for side-by-side savings ($127/year) and access free lab simulators (e.g., 24hr NYC Delivery modules). Prioritize FDA/CLIA-certified courses with seasonal discounts. As outlined in the smart buyer’s guide, combining academic partnerships with hands-on evaluations maximizes ROI while avoiding counterfeit models.